Metabolic reprogramming and tumor angiogenesis are two tightly linked hallmarks of cancer. Regions of high metabolic activity within a tumor can become hypoxic or nutrient starved, eliciting a cascade of pro-angiogenic signals that can lead to increased tumor perfusion and in turn greater metabolic activity. Dynamic contrast enhanced magnetic resonance imaging (DCE-MRI) and 18FDG positron emission tomography (PET) are two imaging methods commonly used in the diagnosis of cancer; the former can be also used to identify the perfusion of the tumor microenvironment (TME) and the latter provides a coarse readout of glucose metabolic activity.
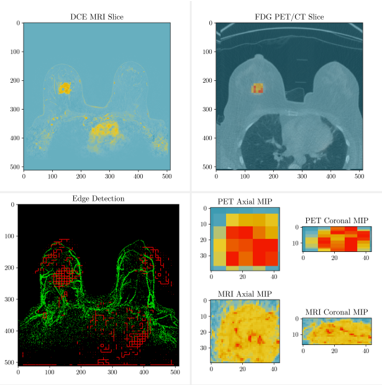
Methods
18 patients received pre-treatment SoC DCE-MRIs, pre-treatment FDG PETs, and 64Cu-DOTA-Trastuzumab PET on the first dose of trastuzumab. We linearly co-registered all images and performed perfusion modeling and multi-tissue segmentation using the DCE-MRI. We then calculated correlations between the PET imaging modalities and perfusion derived parameters in corresponding regions. Finally, we combined perfusion modeling, the tissue segmentation, metabolic modeling, and PKPD modeling to predict response to NACT.
Results
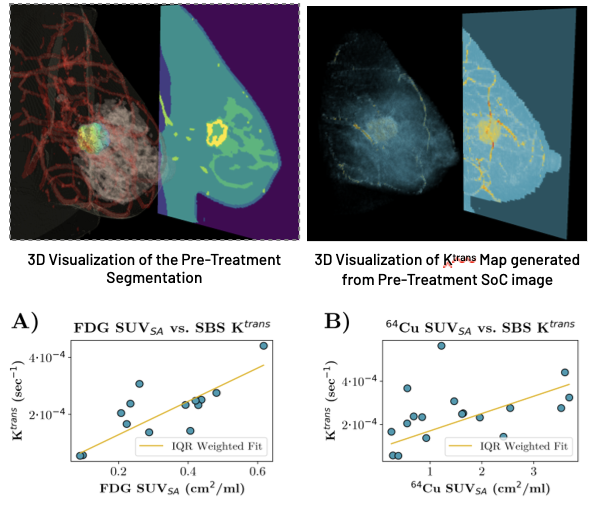
Both PET imaging modalities are shown to correlate well with perfusion parameters; we found that the median SUVSA for both FDG and 64Cu in a tightly cropped region around the lesion was significantly correlated with the median Ktrans generated by SBS-MV from the three-timepoint pre-treatment DCE-MRI.
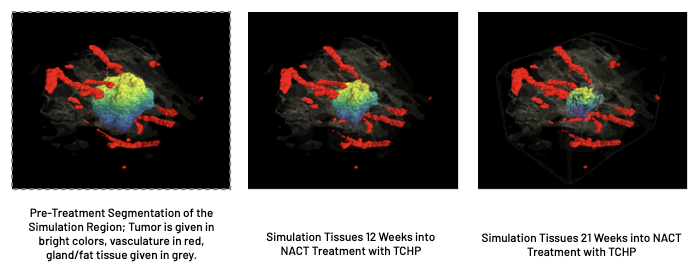
Conclusions
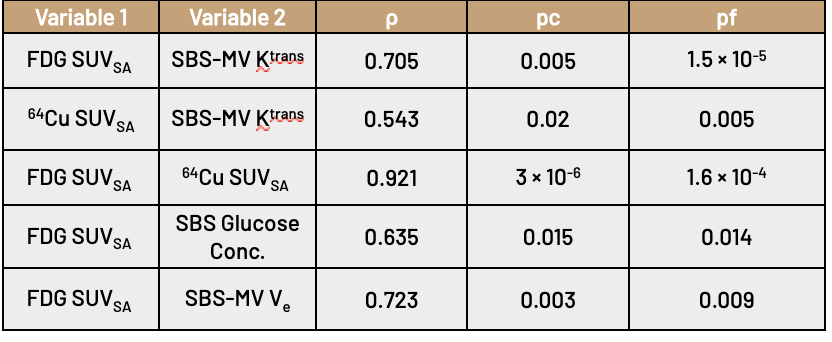
We conclude that our proposed method may be of use significant use in a clinical setting; using only images generated through SoC breast cancer imaging, we constructed perfusion maps and multi-tissue segmentations. Using these maps, we show that the calculated parameters are significantly correlated with PET imaging modalities that are representative of the tumor microenvironment. This result possibly indicates that biomarkers traditionally accessible only through PET imaging may have routes to applicability through current SoC imaging practices.

We also demonstrate that SBS TumorScope is successful in predicting response to NACT in HER2+ patients, performing with an 84% success rate in both predicting pCR and Residual status after treatment at the time of surgery.
Conclusions
Our MV model can extract biologically meaningful perfusion parameters from standard clinical DCE MRI time series, providing the same benefits of a comprehensive kinetic analysis, without impacting current clinical workflow. This approach could be used in both the research and clinical settings, offering actionable information on what drives individual patient therapeutic response for a more personalized care.
About Us
SimBioSys is a software company deploying a combination of biophysical modeling and artificial intelligence to revolutionize precision cancer care. Our portfolio of software applications enables individualized treatment planning, accelerated drug development, clinical trial optimization and comprehensive biomarker development.


